Endonucleases DNA-specific
dsDNase for removal of contaminating DNA from PCR master mixes
Endonucleases DNA-specific
Endonucleases DNA-specific, dsDNase
OverView
Double-Strand Specific dsDNase (dsDNase) is ideal for fast and effective removal of contaminating DNA from PCR master mixes.
Taq polymerases are commonly contaminated by bacterial DNA. This is a problem in PCR based bacterial typing and detection as it might cause false positive results. The unique properties of dsDNase make it suited for removal of contaminating DNA from PCR master mixes prior to addition of DNA template.
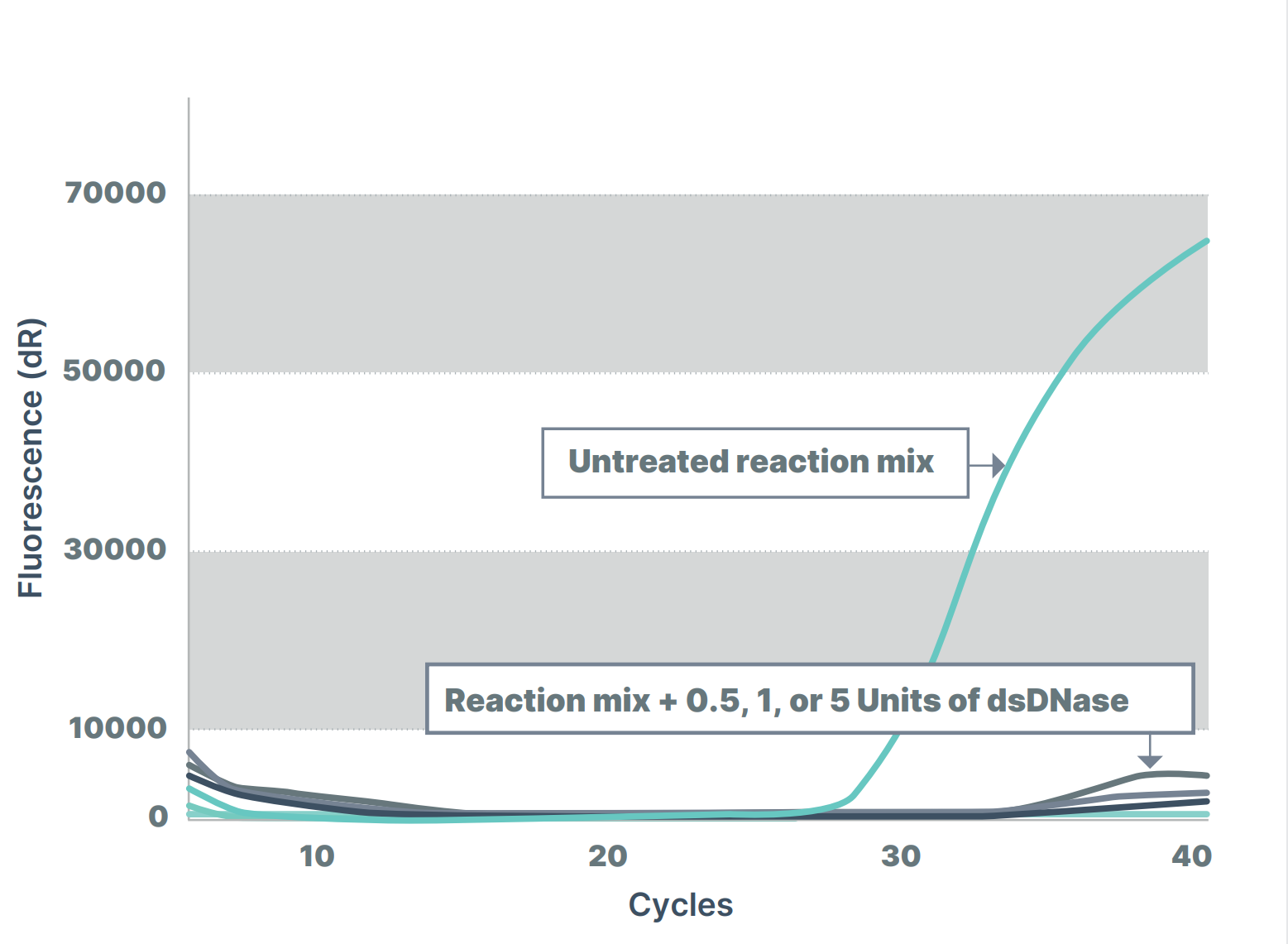
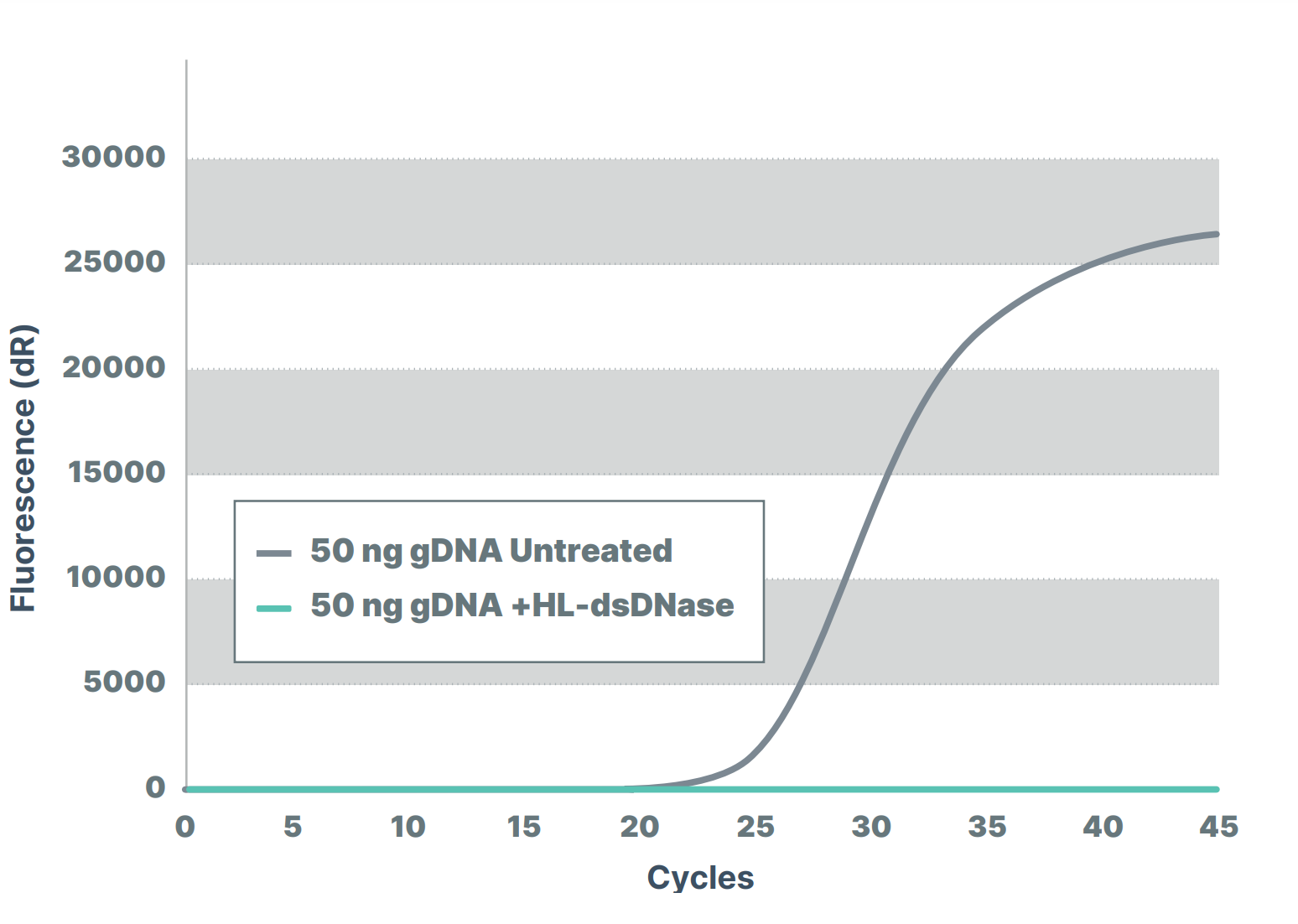
In figure 1, a PCR master mix was treated with different amounts of dsDNase before performing a qPCR to measure the contaminating bacterial DNA in the master mix. ArcticZymes dsDNase effectively removed contaminating DNA below known levels of the assay detection limits.
The dsDNase from Arctic shrimp (Pandalus borealis) is recombinantly produced in Pichia pastoris. It cleaves phosphodiester linkages in DNA to yield oligonucleotides with 5’-phosphate and 3’-hydroxyl termini.
The specific activity is estimated to be 30 times higher than that of bovine DNase I. In the presence of magnesium as only divalent cation and using oligos as a substrate, the activity towards dsDNA is 5000-fold higher than towards ssDNA.
The unique double strand-specificity allows specific degradation of dsDNA while leaving shorter ssDNA as primers and probes essentially intact. Easy inactivation by moderate heat (65°C) allows addition of DNA intended for analysis directly after removal of contaminating DNA.
Workflow – Decontamination of PCR master mixes

Recommended applications:
Key advantages with this Salt Active Nuclease:
- Double-strand DNA specific endonuclease
- High specific activity
- Can be heat-inactivated by moderate heat treatment (65°C for 15 minutes)
- Producing 5′-phospho-oligonucleotide products
Figures
Properties
Specificity towards double-stranded DNA
Nucleic acid specificity has been tested towards double- and single-stranded DNA and RNA oligonucleotides. The specificity of dsDNase towards the substrate has been measured using 15-mer oligonucleotides with FAM at 5′ and DarkQuencher® 3′ (Eurogentec). The fluorescence is proportional to enzyme activity. Assay conditions: 25 mM Tris pH 7.5, 5 mM MgCl2, and 2 μM oligonucleotide.
Relative activities
Substrate Relative Activity
dsDNA 100%
ssDNA <0.03%
dsRNA <0.01%
ssRNA <0.01%
Recommended Protocols
Ordering Info
Documents
Publications
FAQ's
No license required. At ArcticZymes, we pride ourselves on always offering seamless accessibility to our high-quality products. Produced under ISO 13485, our enzymes are sold under a "no license required" policy to ensure that our customer are not restricted by legal burdens, now or with their future use. In addition, we offer our nucleases in a flexible format and are readily available to discuss your customs needs.